USA: Electron Microscopy Why This Research Scratches Your Itch
Scientists at the UNC School of Medicine and the University of California at San Francisco conducted research showing in precise detail how chemicals bind to mast cells to cause an itch. Their work is laying the groundwork for better anti-itching medications with fewer side effects.
Related Vendors
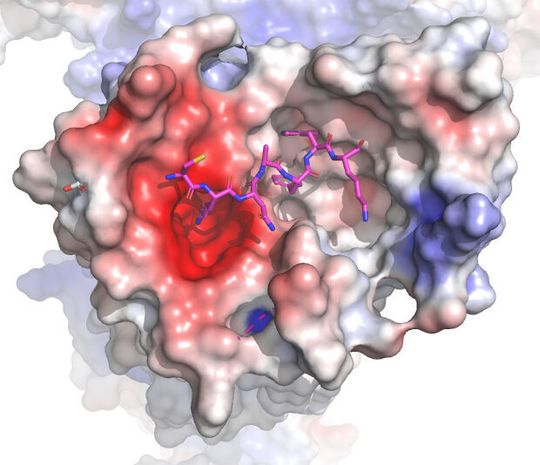
Chapel Hill/USA — UNC-Chapel Hill and UC San Francisco scientists have published work in Nature that provides a template for the design of new anti-itch medication. The scientist showed how chemicals bind to mast cells to cause an itch, and they figured out the detailed structure of receptor proteins on the surface of these cells when a compound is bound to those proteins. This work, published in Nature, was led by the labs of Bryan L. Roth, MD, PhD and Jonathan Fay, PhD at UNC-Chapel Hill, and Brian Shoichet, PhD, at UC San Francisco, co-senior authors who have collaborated on previous studies of important cell receptors — protein complexes that chemicals (including drugs) bind to cause or stop a reaction inside cells. Roth, the Michael Hooker Distinguished Professor of Pharmacology, stated that “..our research team did a truly remarkable job showing precisely how chemically distinct compounds induce itching through one of two distinct receptors known to be involved in itching.”
On the surface of cells sit receptor proteins you can think of as complex locks. When a chemical key enters the lock, not only does the cell “open,” but the chemical causes a chain reaction of signals inside cells. Many chemicals do this, from naturally occurring dopamine in the brain to caffeine and cocaine. When it comes to itch, Roth’s lab identified two receptors called MRGPRX2 on the surface of mast cells and MRGPRX4 on itch-sensing neurons that live in connective tissue and play roles in allergies, immune tolerance, wound healing and other factors in health and disease.
Several drugs unintentionally flood these receptors to trigger the release of histamines, causing the side effect of itching. Drugs such as nateglinide for diabetes, as well as morphine, codeine, and the cough suppressant dextromethorphan are known to cause this reaction. Antihistamines are designed to tamp down the itch response, but they and other anti-itching medications do so clumsily, tripping other cell signaling pathways to cause side effects such as drowsiness, blurred vision, dry mouth, nausea, etc.
The researchers used the experimental technique electron microscopy to create high-resolution maps of these complex receptor proteins when bound to a compound that causes the release of histamines to cause itchiness. They also clarified how drugs bind to MRGPRX4 to cause itch related to various drugs and liver diseases. The researchers used the CryoEM Core Facility at UNC-Chapel Hill to determine the receptor structures.
“Knowing precisely how all this plays out at the molecular level will help us and others create better ways to control the role of these two receptors in itchiness and other conditions,” Roth said. MRGPRX2 and MRGPRX4 have also been implicated in inflammation arising from the nervous system, eczema, ulcerative colitis, and pain. “The relatively potent agonists and antagonists described in our Nature paper provide chemical probes we can use to explore the biology of these receptors,” Roth said, “And the structures we revealed so far should accelerate the search for specific medications targeting MRGPRs.”
References: Structure, function and pharmacology of human itch receptor complexes; Nature, DOI10.1038/s41586-021-04077-y
(ID:47825158)





:quality(80)/p7i.vogel.de/wcms/d6/25/d625408f8628b90bf08a73d0c6d57fe7/0131163821v2.jpeg)
:quality(80)/p7i.vogel.de/wcms/0e/72/0e7296d4ca7d79eb7b3e4f668641af8f/0131148508v2.jpeg)
:quality(80)/p7i.vogel.de/wcms/6b/9a/6b9a2f9d820c458d6518f7d3c0cc990e/0131057985v2.jpeg)
:quality(80)/p7i.vogel.de/wcms/ed/07/ed0756a2495cf3591db2f3c23bedd39c/0131055520v2.jpeg)
:quality(80)/p7i.vogel.de/wcms/52/2e/522e60883334ab2d806e6038a11e9204/0131030069v3.jpeg)
:quality(80)/p7i.vogel.de/wcms/06/0e/060e08234d208d1faca16ad0cf582ff3/0131026470v2.jpeg)
:quality(80)/p7i.vogel.de/wcms/21/6a/216a1b69d48ac4ef7fe694d4c7791531/0131025888v2.jpeg)
:quality(80)/p7i.vogel.de/wcms/4a/8d/4a8d7b8aa03d413a2bbcf34fdf463306/0131025869v2.jpeg)
:quality(80)/p7i.vogel.de/wcms/23/e2/23e2e753cc718480ede0fadd24e0ee02/0131121023v1.jpeg)
:quality(80)/p7i.vogel.de/wcms/b5/63/b5634bbb6ac68f7f55fb261344d6fa1f/0131163645v2.jpeg)
:quality(80)/p7i.vogel.de/wcms/2b/5e/2b5e4798cb0d86b0f29ba4de833c1412/0131003303v1.jpeg)
:quality(80)/p7i.vogel.de/wcms/54/50/5450b302b2cfe1f29c6f3d5c300123ce/0131004604v2.jpeg)
:quality(80)/p7i.vogel.de/wcms/48/97/4897129d96e4ab9287933406b153ab51/0131163956v2.jpeg)
:quality(80)/p7i.vogel.de/wcms/f6/a8/f6a8a2ac333baa08d168cc68ac2192c0/0131001826v2.jpeg)
:quality(80)/p7i.vogel.de/wcms/34/ff/34ff3de563d63ac821f789f8ca397070/0130971132v2.jpeg)
:quality(80)/p7i.vogel.de/wcms/e7/08/e708baaa36f8929e985dfe5baea8bc68/0130734917v2.jpeg)
:quality(80)/p7i.vogel.de/wcms/a2/bf/a2bf01924b12064f93c6f30be4ca6d7a/0131300118v2.jpeg)
:quality(80)/p7i.vogel.de/wcms/e2/fe/e2fee55c4202eaae5c04383233fa8cf3/0131299511v1.jpeg)
:quality(80)/p7i.vogel.de/wcms/a2/a7/a2a7676a95680bba624b1f5b165c8804/0131298919v1.jpeg)
:quality(80)/p7i.vogel.de/wcms/25/d8/25d8dd47da6179f23914adb6bb448757/0131194053v2.jpeg)
:quality(80)/images.vogel.de/vogelonline/bdb/1710600/1710675/original.jpg)
:quality(80)/images.vogel.de/vogelonline/bdb/1682500/1682579/original.jpg)
:quality(80)/images.vogel.de/vogelonline/bdb/1677000/1677080/original.jpg)
:fill(fff,0)/p7i.vogel.de/companies/5f/3f/5f3f63ceedc78/knf-logo.jpg)
:fill(fff,0)/images.vogel.de/vogelonline/companyimg/131900/131910/65.jpg)
:fill(fff,0)/p7i.vogel.de/companies/69/b7/69b7f63f311d9/logo-bimos-tif--2-.jpeg)
:quality(80)/p7i.vogel.de/wcms/0d/9b/0d9bcd7e41846276db862456d619edd2/0129351002v2.jpeg)
:quality(80)/p7i.vogel.de/wcms/66/5a/665af659d8aa6adaf1eedece354cecc9/0125742135v2.jpeg)