Germany: Molecular Biology Setup and Start within a Few Seconds: How Cells Choose Active Genes Correctly
It is essential for cells to control precisely which of the many genes of their genetic material they use. This is done in so-called transcription factories, molecular clusters in the nucleus. Researchers have now found that the formation of transcription factories resembles the condensation of liquids.
Related Vendors
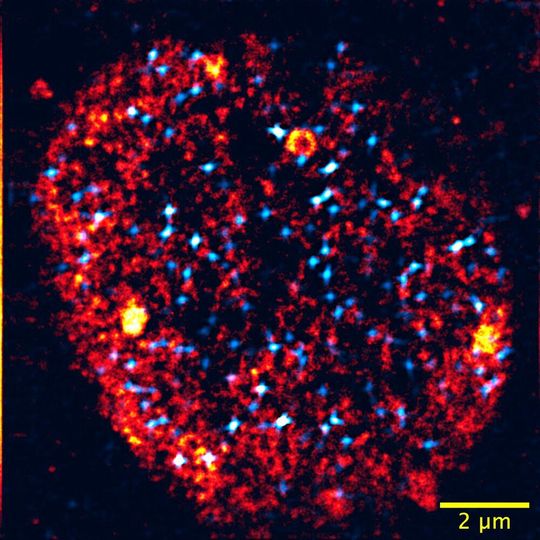
Karlsruhe/Germany — Human genetic material contains more than 20,000 different genes. But each cell only uses a fraction of the information stored in this genome. Hence, cells have to control precisely which genes they use. If not, cancer or embryonal growth disorder may develop. So-called transcription factories play a central role in the selection of active genes. “These factories are molecular clusters in the nucleus that combine the correct selection of active genes and the read-out of their sequence at a central location,” Lennart Hilbert explains. The Junior Professor for Systems Biology/Bioinformatics at the Zoological Institute (ZOO) of KIT also heads a working group at KIT’s Institute of Biological and Chemical Systems — Biological Information Processing (IBCS-BIP).
For decades, cellular and molecular biologists have studied how transcription factories are set up and taken into operation within a few seconds. Results obtained so far suggest relevance of processes known from industrial and technical polymer and liquid materials only. Current research focuses on phase separation as a central mechanism. In everyday life, phase separation can be observed when separating oil from water. It has not yet been clear, however, how exactly phase separation contributes to the setup of transcription factories in living cells.
Researchers from KIT’s Institute of Biological and Chemical Systems (IBCS), Zoological Institute (ZOO), Institute of Applied Physics (APH), and Institute of Nanotechnology (INT), in cooperation with scientists from FAU and MPZPM in Erlangen and the University of Illinois at Urbana-Champaign/USA, have now gained new findings on the formation of transcription factories: It is similar to the condensation of liquids. This is reported in Molecular Systems Biology. The first co-authors are Agnieszka Pancholi of IBCS-BIP and ZOO and Tim Klingberg of FAU and MPZPM.
Latest Light Microscopy Combined with Computer Simulations
In their publication, the researchers point out that condensation to form transcription factories resembles steamy glasses or windows. Liquid condenses in the presence of a receptive surface only, but then very quickly. In the living cell, specially marked areas of the genome are used as condensation surfaces. The liquid-coated areas allow for the adhesion of relevant gene sequences and additional molecules that eventually activate the adhering genes. These findings were obtained by interdisciplinary cooperation. Zebrafish embryos were studied with latest light microscopes developed by Professor Gerd Ulrich Nienhaus’s Chair at APH. These observations were then linked to computer simulations at the FAU Chair for Mathematics headed by Professor Vasily Zaburdaev. Combination of observations and simulations makes the condensation process reproducible and explains how living cells can set up transcription factories rapidly and reliably.
New understanding of condensed liquids in living cells recently resulted in entirely new approaches to treating cancer and diseases of the nervous system. These approaches are now being pursued by startups developing new drugs. Other research activities focus on the use of DNA sequences as digital data storage systems. Meanwhile, principle feasibility of DNA-based data storage systems has been demonstrated by several working groups. Reliable storage and read-out of information in such DNA storage media still represent big challenges. “Our work shows how the biological cell organizes such processes rapidly and reliably. The computer simulations and functional concepts developed by us can be transferred directly to artificial DNA systems and can support their design,” Lennart Hilbert says.
Original Publication: Agnieszka Pancholi, Tim Klingberg, Weichun Zhang, Roshan Prizak, Irina Mamontova, Amra Noa, Marcel Sobucki, Andrei Yu Kobitski, Gerd Ulrich Nienhaus, Vasily Zaburdaev, Lennart Hilbert: RNA polymerase II clusters form in line with surface condensation on regulatory chromatin. Molecular Systems Biology, 2021. DOI: 10.15252/msb.202110272
(ID:47779113)





:quality(80)/p7i.vogel.de/wcms/6b/9a/6b9a2f9d820c458d6518f7d3c0cc990e/0131057985v2.jpeg)
:quality(80)/p7i.vogel.de/wcms/ed/07/ed0756a2495cf3591db2f3c23bedd39c/0131055520v2.jpeg)
:quality(80)/p7i.vogel.de/wcms/f1/87/f187c86e791dfcf8ecccf615a6a3d054/0131055400v2.jpeg)
:quality(80)/p7i.vogel.de/wcms/cb/39/cb3902f5777a11e07655ba29d00a0394/0131003637v1.jpeg)
:quality(80)/p7i.vogel.de/wcms/52/2e/522e60883334ab2d806e6038a11e9204/0131030069v3.jpeg)
:quality(80)/p7i.vogel.de/wcms/06/0e/060e08234d208d1faca16ad0cf582ff3/0131026470v2.jpeg)
:quality(80)/p7i.vogel.de/wcms/21/6a/216a1b69d48ac4ef7fe694d4c7791531/0131025888v2.jpeg)
:quality(80)/p7i.vogel.de/wcms/4a/8d/4a8d7b8aa03d413a2bbcf34fdf463306/0131025869v2.jpeg)
:quality(80)/p7i.vogel.de/wcms/2b/5e/2b5e4798cb0d86b0f29ba4de833c1412/0131003303v1.jpeg)
:quality(80)/p7i.vogel.de/wcms/54/50/5450b302b2cfe1f29c6f3d5c300123ce/0131004604v2.jpeg)
:quality(80)/p7i.vogel.de/wcms/f1/71/f1711b934d493949c55d6e9b9ddcffe9/0131003077v1.jpeg)
:quality(80)/p7i.vogel.de/wcms/60/54/6054287023c79b93d166c8b7802f9ad4/0130972957v2.jpeg)
:quality(80)/p7i.vogel.de/wcms/f6/a8/f6a8a2ac333baa08d168cc68ac2192c0/0131001826v2.jpeg)
:quality(80)/p7i.vogel.de/wcms/34/ff/34ff3de563d63ac821f789f8ca397070/0130971132v2.jpeg)
:quality(80)/p7i.vogel.de/wcms/e7/08/e708baaa36f8929e985dfe5baea8bc68/0130734917v2.jpeg)
:quality(80)/p7i.vogel.de/wcms/ea/5a/ea5a5ed15b2c98730c7111bb56b1a7aa/0130612537v1.jpeg)
:quality(80)/p7i.vogel.de/wcms/33/50/3350efd72f2c620af7c2133ce55c901f/0131086519v2.jpeg)
:quality(80)/p7i.vogel.de/wcms/ca/fa/cafafb635ff66815b96824c2f099968a/0131058329v2.jpeg)
:quality(80)/p7i.vogel.de/wcms/56/d7/56d73ff8d8ffc3ce50e3ff2188b7aada/0131004724v2.jpeg)
:quality(80)/p7i.vogel.de/wcms/87/d5/87d5890059d61e8107148c2039ff9bd8/0131004713v2.jpeg)
:quality(80)/images.vogel.de/vogelonline/bdb/1710600/1710675/original.jpg)
:quality(80)/images.vogel.de/vogelonline/bdb/1682500/1682579/original.jpg)
:quality(80)/images.vogel.de/vogelonline/bdb/1677000/1677080/original.jpg)
:fill(fff,0)/p7i.vogel.de/companies/5f/3f/5f3f63ceedc78/knf-logo.jpg)
:fill(fff,0)/images.vogel.de/vogelonline/companyimg/131900/131910/65.jpg)
:fill(fff,0)/p7i.vogel.de/companies/69/b7/69b7f63f311d9/logo-bimos-tif--2-.jpeg)
:quality(80)/p7i.vogel.de/wcms/ed/07/ed0756a2495cf3591db2f3c23bedd39c/0131055520v2.jpeg)
:quality(80)/p7i.vogel.de/wcms/e2/83/e28330d9bc97092bbdc4d7d61b2103f5/0125079999v2.jpeg)